© Shipunov A.B. Systema Naturae [Electronic resource]. 1995—onwards.
Mode of
access: http://herba.msu.ru/shipunov/os/os-en.htm
Systema Naturae
Current version:
Publications and older versions:
This is the original variant of the classification of living organisms. The classification has: (1) small number of kingdoms; (2) large phyla; (3) incorporating as much as possible taxonomical data from different sources. Please intend that order of taxa unlike usual practice, is important alongside with hierarchical structure.
Short description
- The classification utilizes all described groups of living organisms, excluding fossils and viruses, from kingdoms and down to the level of class. Some properly described "environmental DNA" groups are also included.
- Initial fragmentation of taxonomic space is based on pre-existed hypotheses of how groups should be defined: "levels of complexity" for kingdoms (see below), "structure of life cycle" for plants, "bauplan" for animals. In cases where these hypotheses are absent, the most stable and inclusive molecular groups corrected with morphological evidence are used.
- Subsequent (and iterative) fragmentation of taxonomic space based on distance approach. Distances are obtained from "total evidence" of all available and up-to-date primary (like morphological or single molecular characters) and secondary characters (suggestions of similarity taken from different sources, e.g., trees). All characters were weighted.
- Since the problem of paraphyly is in the first place the problem of communication between tree-based and space-based approaches, some paraphyletic groups are accepted, generally in cases of (a) unresolved, unstable and/or non-typical (like nets) structures in trees; (b) high inequality/irregularity in trees; (c) dense and multiply ("comb-like") grades.
- The final criteria for the classification are (a) simplicity, (b) predictability and (c) evolutionary/ecological integrity.
- All groups treated in the most inclusive manner.
- Classification is fully hierarchical; ranks should be comparable, at least across the largest divisions (kingdoms). The order of taxa is meaningful and reflects the position of taxon in the taxonomic space.
About kingdoms
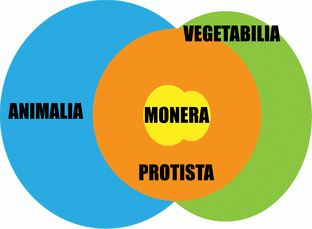
Kingdoms are the highest available subdivisions of the diversity. Four kingdoms are accepted: Monera, Protista, Vegetabilia and Animalia, which represent different steps of increased complexity of living things, from simple prokaryotic cell to compound eukaryotic cell and further to tissue/organ cell systems.
Typified names
Since most so-called "trivial" names for higher categories are used in very different senses, and often dubious, this classification introduces the use of globally typified names with rank designation. Such a name is constructed from the name of type genus and rank designation (superscript number at the upper left on the name), like 5Name. I beleive that these numberical ranks are better than traditional endings, which usually mask the basionym, hard to remember, sometimes conflicting (for example, among protists where different nomenclature codes work together) and looks non-aesthetic.
Each basic rank have the number:
| Species | 1
|
| Genus | 2
|
| Familia | 3
|
| Order | 4
|
| Classis | 5
|
| Phylum/Divisio | 6
|
| Regnum | 7
|
Intermediate ranges are designated as decimal numbers:
| Sub- | 0.8
|
| Infra- | 0.5
|
| Super- | 0.2
|
Thus, 5.2Name means this is a taxon with rank of superclass based on genus "Name". I hope that it is quite simple and efficient system.
dactylorhiza@gmail.com
Back

